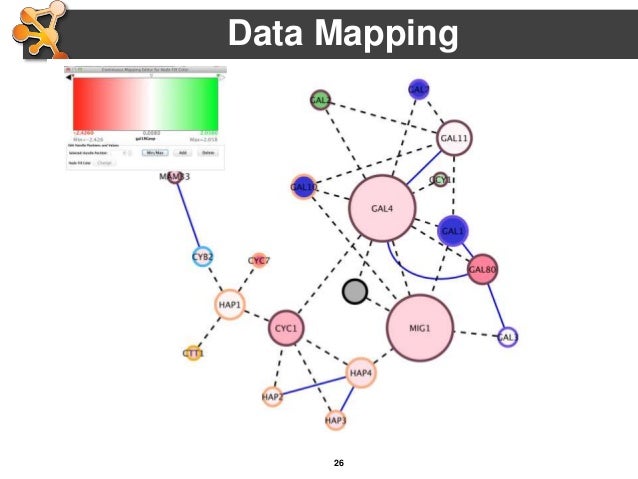
Physical Interaction: Protein-protein interaction data.Most of these data are collected from the Gene Expression Omnibus (GEO) we only collect data associated with a publication. Two genes are linked if their expression levels are similar across conditions in a gene expression study. Networks names describe the data source and are either generated from the PubMed entry associated with the data source (first author-last author-year), or simply the name of the data source (BioGRID, Pathwa圜ommons-(original data source), Pfam) These include protein-protein, protein-DNA and genetic interactions, pathways, reactions, gene and protein expression data, protein domains and phenotypic screening profiles. GeneMANIA searches many large, publicly available biological datasets to find related genes. Access the advanced options by clicking the ellipsis (“…”) in the search bar. The number of resultant genes, the number of resultant attributes, and the weighting method can be configured in the advanced options.The GeneMANIA Cytoscape app is capable of handling larger gene lists. 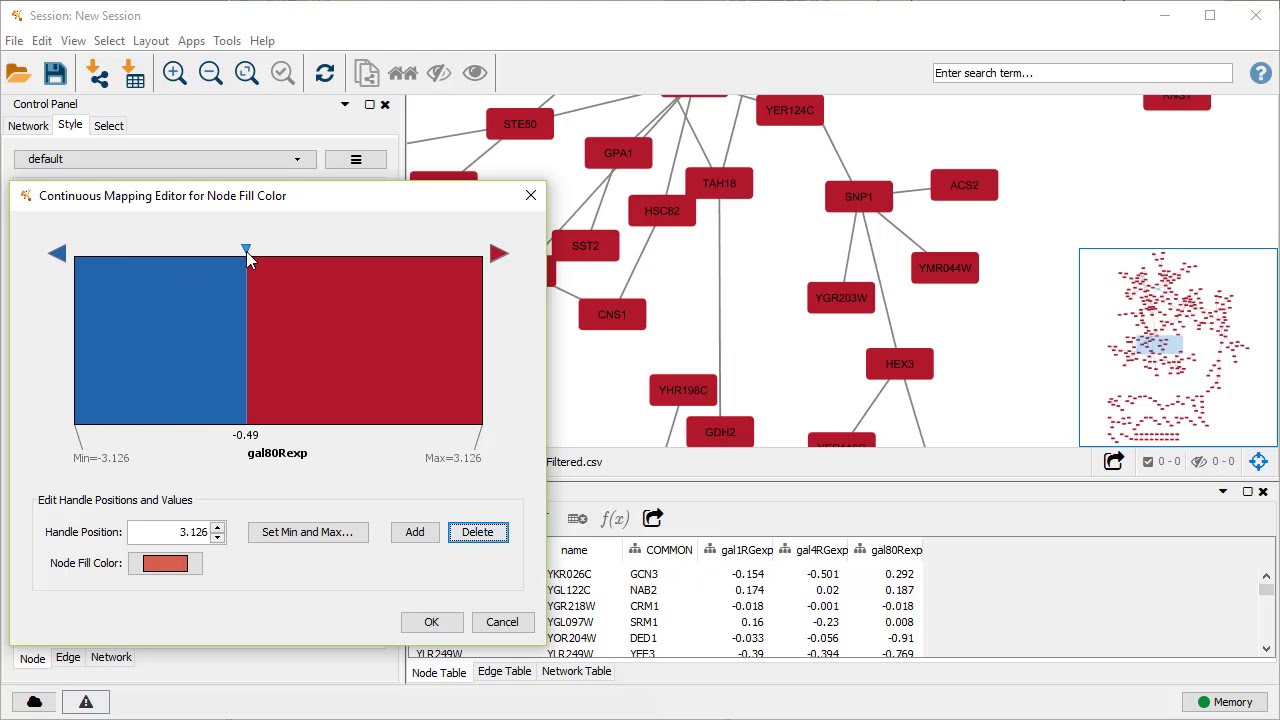
GeneMANIA will be slower with an input gene list of more than 50 genes if you have such large gene lists, we recommend using a gene list of no more than 100 genes.If your query list has less than 6 genes, GeneMANIA will make gene function predictions based on GO annotations patterns. If your query list consists of 6 or more genes, GeneMANIA will calculate gene list-specific weights.It does not matter which function they are related by, as long as that function is captured somehow by some functional association networks in the GeneMANIA system. If they are not, a disconnected network will result and the network weighting will not be optimal. GeneMANIA works best if most of the input genes are functionally related.Choosing an appropriate network weighting option.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
